(P18) Towards HIV diagnostics at the point-of-care using rolling circle amplification and microfluidic signal enhancement
Författare/Medförfattare
Ruben R. G. Soares [1], Sibel Ciftci [2], João C. Varela [2], Ashokkumar Manickam [3], Ujjwal Neogi [3,4], Mats Nilsson [2], Narayanan Madaboosi [2], Aman Russom [1]
Affiliates
[1] KTH Royal Institute of Technology, Science for Life Laboratory, Solna, SWEDEN [2] Stockholm University, Science for Life Laboratory, Solna, SWEDEN [3] Karolinska Institutet, Stockholm, SWEDEN [4] University of Missouri, Columbia, USA
Abstract
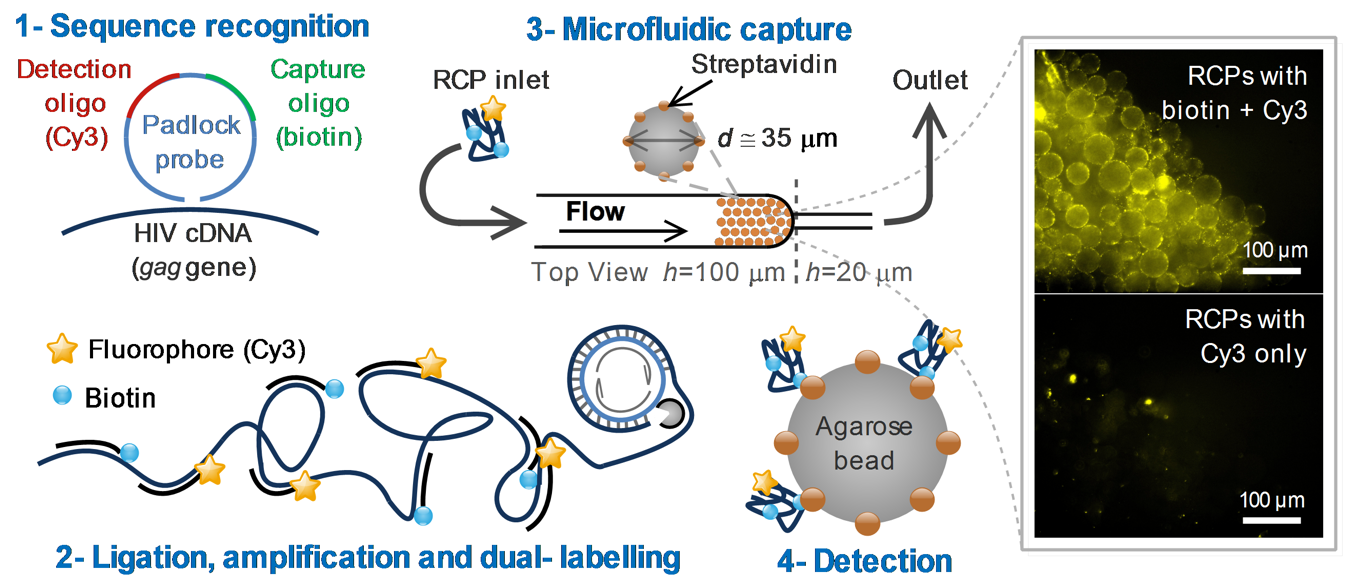
HIV infections are devastating in resource-limited settings, where portable and affordable diagnostic devices would aid effective monitoring of infection spread as well as therapeutics. To develop a cost-effective, specific and sensitive biosensor for HIV diagnostics, isothermal molecular assays like Rolling Circle Amplification (RCA) are desirable to minimize technical complexities involved with rapid temperature ramping. However, multiple rounds of RCA are typically required to detect clinically relevant virus titers in samples, implying longer assays and associated technical complexities. To improve the assay yield, the spatial concentration of RCA products (RCPs), typically read as discrete digital signals, is critical to boost the sensitivity using a single round of RCA. Here, an RCA-based assay using multiple padlock probes (PLPs) targeting the conserved gag gene of HIV-1 subtype B is presented. The detection of HIV load was performed in a microfluidic device exploiting a dual-labeling strategy, using biotin and fluorophore, for signal amplification.
The RCA was performed for 120 min at 37 °C using ϕ29 polymerase with 10 µL of sample. The RCPs were labeled using two detection oligos labeled with either Cy3 or biotin, both complementary to specific regions in the PLP backbone. The RCPs were captured in a PDMS microchannel by flowing the solution directly through packed streptavidin-conjugated beads (~ 100 nL) at 2.5 µL/min for 15 min, followed by fluorescence microscopy imaging.
The microfluidic readout, coupled to RCA, resulted in detecting down to 10 fM of synthetic target, i.e. 0.1 amol ssDNA in the sample. This value is approximately 100-fold lower compared to conventional RCP counting on glass slides and > 20-fold lower compared to a recently published silica bead-based microfluidic RCP capture [1]. The validated microfluidic detection platform was further used to measure RCPs generated using HIV-1 pNL4-3 viral RNA, obtained by transfecting 293T cell line and collecting the supernatant after 3 days. Reverse transcription (SuperScript III reverse transcriptase) was performed on the samples prior to RCA. After measuring 3 serial dilutions (2-,10- and 50-fold) of the initial supernatant of infected cells (65.2 ng/µL total RNA) and a 2-fold diluted supernatant from non-infected cells, the results highlight a direct correlation between number of counted RCPs and the initial RNA concentration, as well as negligible non-specific signal provided by the non-infected cell culture supernatant. Finally, the device was used to successfully measure 11 NIH standards of HIV subtypes B and C (P24 concentration ranging from ~3 up to ~127 ng/mL), as well as circulating recombinant forms CRF02_AE and CRF02_AG, significantly above the background provided by the non-infected cell culture supernatant.
These findings highlight the compatibility of the developed set of PLPs with the detection of several HIV subtypes and the enhancement in sensitivity provided by microfluidics when combined with the dual-labeling strategy, thus paving the way to achieve clinical relevance.
[1] R. R. G. Soares et al, Biosens. Bioelectron. 2019, 128, 68-75